tst = CrossEntropyLossFlat(reduction='none')
output = torch.randn(32, 5, 10)
target = torch.randint(0, 10, (32,5))
#nn.CrossEntropy would fail with those two tensors, but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.CrossEntropyLoss()(output,target))
#Associated activation is softmax
test_eq(tst.activation(output), F.softmax(output, dim=-1))
#This loss function has a decodes which is argmax
test_eq(tst.decodes(output), output.argmax(dim=-1))Loss Functions
BaseLoss
def BaseLoss(
loss_cls, # Uninitialized PyTorch-compatible loss
args:VAR_POSITIONAL, axis:int=-1, # Class axis
flatten:bool=True, # Flatten `inp` and `targ` before calculating loss
floatify:bool=False, # Convert `targ` to `float`
is_2d:bool=True, # Whether `flatten` keeps one or two channels when applied
kwargs:VAR_KEYWORD
):
Same as loss_cls, but flattens input and target.
Wrapping a general loss function inside of BaseLoss provides extra functionalities to your loss functions:
- flattens the tensors before trying to take the losses since it’s more convenient (with a potential tranpose to put
axisat the end) - a potential
activationmethod that tells the library if there is an activation fused in the loss (useful for inference and methods such asLearner.get_predsorLearner.predict) - a potential
decodesmethod that is used on predictions in inference (for instance, an argmax in classification)
The args and kwargs will be passed to loss_cls during the initialization to instantiate a loss function. axis is put at the end for losses like softmax that are often performed on the last axis. If floatify=True, the targs will be converted to floats (useful for losses that only accept float targets like BCEWithLogitsLoss), and is_2d determines if we flatten while keeping the first dimension (batch size) or completely flatten the input. We want the first for losses like Cross Entropy, and the second for pretty much anything else.
CrossEntropyLossFlat
def CrossEntropyLossFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
weight:NoneType=None, ignore_index:int=-100, reduction:str='mean',
flatten:bool=True, # Flatten `inp` and `targ` before calculating loss
floatify:bool=False, # Convert `targ` to `float`
is_2d:bool=True, # Whether `flatten` keeps one or two channels when applied
):
Same as nn.CrossEntropyLoss, but flattens input and target.
#In a segmentation task, we want to take the softmax over the channel dimension
tst = CrossEntropyLossFlat(axis=1)
output = torch.randn(32, 5, 128, 128)
target = torch.randint(0, 5, (32, 128, 128))
_ = tst(output, target)
test_eq(tst.activation(output), F.softmax(output, dim=1))
test_eq(tst.decodes(output), output.argmax(dim=1))Focal Loss is the same as cross entropy except easy-to-classify observations are down-weighted in the loss calculation. The strength of down-weighting is proportional to the size of the gamma parameter. Put another way, the larger gamma the less the easy-to-classify observations contribute to the loss.
FocalLossFlat
def FocalLossFlat(
args:VAR_POSITIONAL,
gamma:float=2.0, # Focusing parameter. Higher values down-weight easy examples' contribution to loss
axis:int=-1, # Class axis
weight:NoneType=None, reduction:str='mean', kwargs:VAR_KEYWORD
):
Same as CrossEntropyLossFlat but with focal paramter, gamma. Focal loss is introduced by Lin et al. https://arxiv.org/pdf/1708.02002.pdf. Note the class weighting factor in the paper, alpha, can be implemented through pytorch weight argument passed through to F.cross_entropy.
FocalLoss
def FocalLoss(
gamma:float=2.0, # Focusing parameter. Higher values down-weight easy examples' contribution to loss
weight:Tensor=None, # Manual rescaling weight given to each class
reduction:str='mean', # PyTorch reduction to apply to the output
):
Same as nn.Module, but no need for subclasses to call super().__init__
#Compare focal loss with gamma = 0 to cross entropy
fl = FocalLossFlat(gamma=0)
ce = CrossEntropyLossFlat()
output = torch.randn(32, 5, 10)
target = torch.randint(0, 10, (32,5))
test_close(fl(output, target), ce(output, target))
#Test focal loss with gamma > 0 is different than cross entropy
fl = FocalLossFlat(gamma=2)
test_ne(fl(output, target), ce(output, target))#In a segmentation task, we want to take the softmax over the channel dimension
fl = FocalLossFlat(gamma=0, axis=1)
ce = CrossEntropyLossFlat(axis=1)
output = torch.randn(32, 5, 128, 128)
target = torch.randint(0, 5, (32, 128, 128))
test_close(fl(output, target), ce(output, target), eps=1e-4)
test_eq(fl.activation(output), F.softmax(output, dim=1))
test_eq(fl.decodes(output), output.argmax(dim=1))BCEWithLogitsLossFlat
def BCEWithLogitsLossFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
floatify:bool=True, # Convert `targ` to `float`
thresh:float=0.5, # The threshold on which to predict
weight:NoneType=None, reduction:str='mean', pos_weight:NoneType=None,
flatten:bool=True, # Flatten `inp` and `targ` before calculating loss
is_2d:bool=True, # Whether `flatten` keeps one or two channels when applied
):
Same as nn.BCEWithLogitsLoss, but flattens input and target.
tst = BCEWithLogitsLossFlat()
output = torch.randn(32, 5, 10)
target = torch.randn(32, 5, 10)
#nn.BCEWithLogitsLoss would fail with those two tensors, but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.BCEWithLogitsLoss()(output,target))
output = torch.randn(32, 5)
target = torch.randint(0,2,(32, 5))
#nn.BCEWithLogitsLoss would fail with int targets but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.BCEWithLogitsLoss()(output,target))
tst = BCEWithLogitsLossFlat(pos_weight=torch.ones(10))
output = torch.randn(32, 5, 10)
target = torch.randn(32, 5, 10)
_ = tst(output, target)
test_fail(lambda x: nn.BCEWithLogitsLoss()(output,target))
#Associated activation is sigmoid
test_eq(tst.activation(output), torch.sigmoid(output))BCELossFlat
def BCELossFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
floatify:bool=True, # Convert `targ` to `float`
weight:NoneType=None, reduction:str='mean'
):
Same as nn.BCELoss, but flattens input and target.
tst = BCELossFlat()
output = torch.sigmoid(torch.randn(32, 5, 10))
target = torch.randint(0,2,(32, 5, 10))
_ = tst(output, target)
test_fail(lambda x: nn.BCELoss()(output,target))MSELossFlat
def MSELossFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
floatify:bool=True, # Convert `targ` to `float`
reduction:str='mean'
):
Same as nn.MSELoss, but flattens input and target.
tst = MSELossFlat()
output = torch.sigmoid(torch.randn(32, 5, 10))
target = torch.randint(0,2,(32, 5, 10))
_ = tst(output, target)
test_fail(lambda x: nn.MSELoss()(output,target))L1LossFlat
def L1LossFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
floatify:bool=True, # Convert `targ` to `float`
reduction:str='mean'
):
Same as nn.L1Loss, but flattens input and target.
LabelSmoothingCrossEntropy
def LabelSmoothingCrossEntropy(
eps:float=0.1, # The weight for the interpolation formula
weight:Tensor=None, # Manual rescaling weight given to each class passed to `F.nll_loss`
reduction:str='mean', # PyTorch reduction to apply to the output
):
Same as nn.Module, but no need for subclasses to call super().__init__
lmce = LabelSmoothingCrossEntropy()
output = torch.randn(32, 5, 10)
target = torch.randint(0, 10, (32,5))
test_close(lmce(output.flatten(0,1), target.flatten()), lmce(output.transpose(-1,-2), target))On top of the formula we define:
- a
reductionattribute, that will be used when we callLearner.get_preds weightattribute to pass to BCE.- an
activationfunction that represents the activation fused in the loss (since we use cross entropy behind the scenes). It will be applied to the output of the model when callingLearner.get_predsorLearner.predict - a
decodesfunction that converts the output of the model to a format similar to the target (here indices). This is used inLearner.predictandLearner.show_resultsto decode the predictions
LabelSmoothingCrossEntropyFlat
def LabelSmoothingCrossEntropyFlat(
args:VAR_POSITIONAL, axis:int=-1, # Class axis
eps:float=0.1, reduction:str='mean', flatten:bool=True, # Flatten `inp` and `targ` before calculating loss
floatify:bool=False, # Convert `targ` to `float`
is_2d:bool=True, # Whether `flatten` keeps one or two channels when applied
):
Same as LabelSmoothingCrossEntropy, but flattens input and target.
#These two should always equal each other since the Flat version is just passing data through
lmce = LabelSmoothingCrossEntropy()
lmce_flat = LabelSmoothingCrossEntropyFlat()
output = torch.randn(32, 5, 10)
target = torch.randint(0, 10, (32,5))
test_close(lmce(output.transpose(-1,-2), target), lmce_flat(output,target))We present a general Dice loss for segmentation tasks. It is commonly used together with CrossEntropyLoss or FocalLoss in kaggle competitions. This is very similar to the DiceMulti metric, but to be able to derivate through, we replace the argmax activation by a softmax and compare this with a one-hot encoded target mask. This function also adds a smooth parameter to help numerical stabilities in the intersection over union division. If your network has problem learning with this DiceLoss, try to set the square_in_union parameter in the DiceLoss constructor to True.
DiceLoss
def DiceLoss(
axis:int=1, # Class axis
smooth:float=1e-06, # Helps with numerical stabilities in the IoU division
reduction:str='sum', # PyTorch reduction to apply to the output
square_in_union:bool=False, # Squares predictions to increase slope of gradients
):
Dice loss for segmentation
dl = DiceLoss()
_x = tensor( [[[1, 0, 2],
[2, 2, 1]]])
_one_hot_x = tensor([[[[0, 1, 0],
[0, 0, 0]],
[[1, 0, 0],
[0, 0, 1]],
[[0, 0, 1],
[1, 1, 0]]]])
test_eq(dl._one_hot(_x, 3), _one_hot_x)dl = DiceLoss()
model_output = tensor([[[[2., 1.],
[1., 5.]],
[[1, 2.],
[3., 1.]],
[[3., 0],
[4., 3.]]]])
target = tensor([[[2, 1],
[2, 0]]])
dl_out = dl(model_output, target)
test_eq(dl.decodes(model_output), target)dl = DiceLoss(reduction="mean")
#identical masks
model_output = tensor([[[.1], [.1], [100.]]])
target = tensor([[2]])
test_close(dl(model_output, target), 0)
#50% intersection
model_output = tensor([[[.1, 100.], [.1, .1], [100., .1]]])
target = tensor([[2, 1]])
test_close(dl(model_output, target), .66, eps=0.01)As a test case for the dice loss consider satellite image segmentation. Let us say we have three classes: Background (0), River (1) and Road (2). Let us look at a specific target
target = torch.zeros(100,100)
target[:,5] = 1
target[:,50] = 2
plt.imshow(target);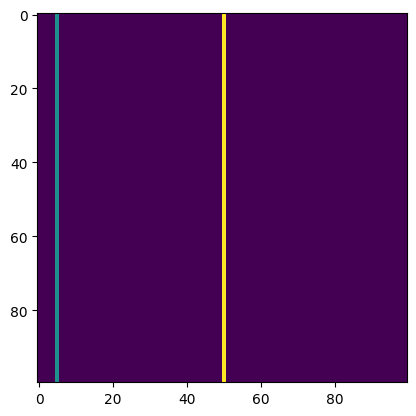
Nearly everything is background in this example, and we have a thin river at the left of the image as well as a thin road in the middle of the image. If all our data looks similar to this, we say that there is a class imbalance, meaning that some classes (like river and road) appear relatively infrequently. If our model just predicted “background” (i.e. the value 0) for all pixels, it would be correct for most pixels. But this would be a bad model and the diceloss should reflect that
model_output_all_background = torch.zeros(3, 100,100)
# assign probability 1 to class 0 everywhere
# to get probability 1, we just need a high model output before softmax gets applied
model_output_all_background[0,:,:] = 100# add a batch dimension
model_output_all_background = torch.unsqueeze(model_output_all_background,0)
target = torch.unsqueeze(target,0)Our dice score should be around 1/3 here, because the “background” class is predicted correctly (and that for nearly every pixel), but the other two clases are never predicted correctly. Dice score of 1/3 means dice loss of 1 - 1/3 = 2/3:
test_close(dl(model_output_all_background, target), 0.67, eps=0.01)If the model would predict everything correctly, the dice loss should be zero:
correct_model_output = torch.zeros(3, 100,100)
correct_model_output[0,:,:] = 100
correct_model_output[0,:,5] = 0
correct_model_output[0,:,50] = 0
correct_model_output[1,:,5] = 100
correct_model_output[2,:,50] = 100
correct_model_output = torch.unsqueeze(correct_model_output, 0)test_close(dl(correct_model_output, target), 0)You could easily combine this loss with FocalLoss defining a CombinedLoss, to balance between global (Dice) and local (Focal) features on the target mask.
class CombinedLoss:
"Dice and Focal combined"
def __init__(self, axis=1, smooth=1., alpha=1.):
store_attr()
self.focal_loss = FocalLossFlat(axis=axis)
self.dice_loss = DiceLoss(axis, smooth)
def __call__(self, pred, targ):
return self.focal_loss(pred, targ) + self.alpha * self.dice_loss(pred, targ)
def decodes(self, x): return x.argmax(dim=self.axis)
def activation(self, x): return F.softmax(x, dim=self.axis)cl = CombinedLoss()
output = torch.randn(32, 4, 5, 10)
target = torch.randint(0,2,(32, 5, 10))
_ = cl(output, target)# Tests to catch future changes to pickle which cause some loss functions to be 'unpicklable'.
# This causes problems with `Learner.export` as the model can't be pickled with these particular loss funcitons.
losses_picklable = [
(BCELossFlat(), True),
(BCEWithLogitsLossFlat(), True),
(CombinedLoss(), True),
(CrossEntropyLossFlat(), True),
(DiceLoss(), True),
(FocalLoss(), True),
(FocalLossFlat(), True),
(L1LossFlat(), True),
(LabelSmoothingCrossEntropyFlat(), True),
(LabelSmoothingCrossEntropy(), True),
(MSELossFlat(), True),
]
for loss, picklable in losses_picklable:
try:
pickle.dumps(loss, protocol=2)
except (pickle.PicklingError, TypeError) as e:
if picklable:
# Loss was previously picklable but isn't currently
raise e